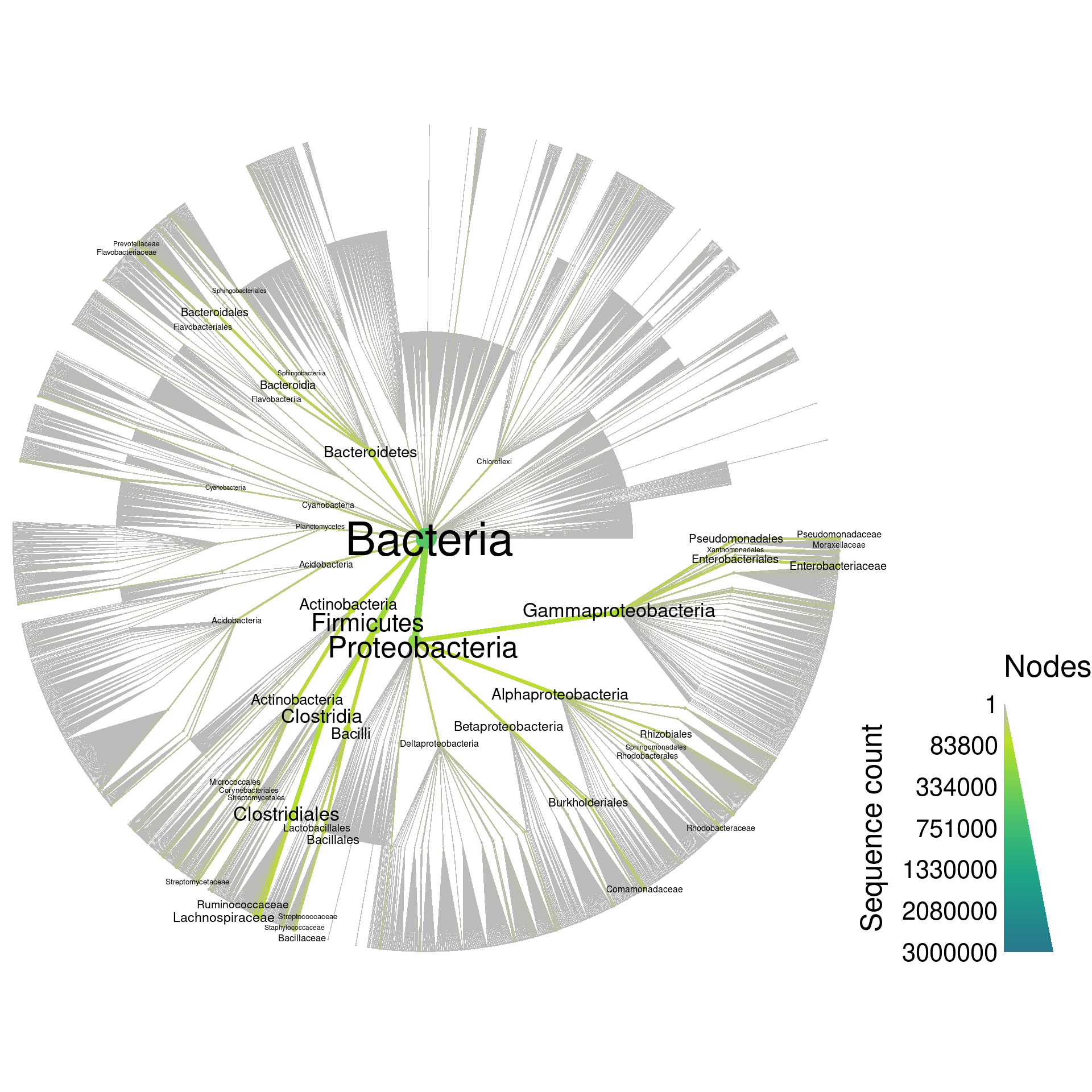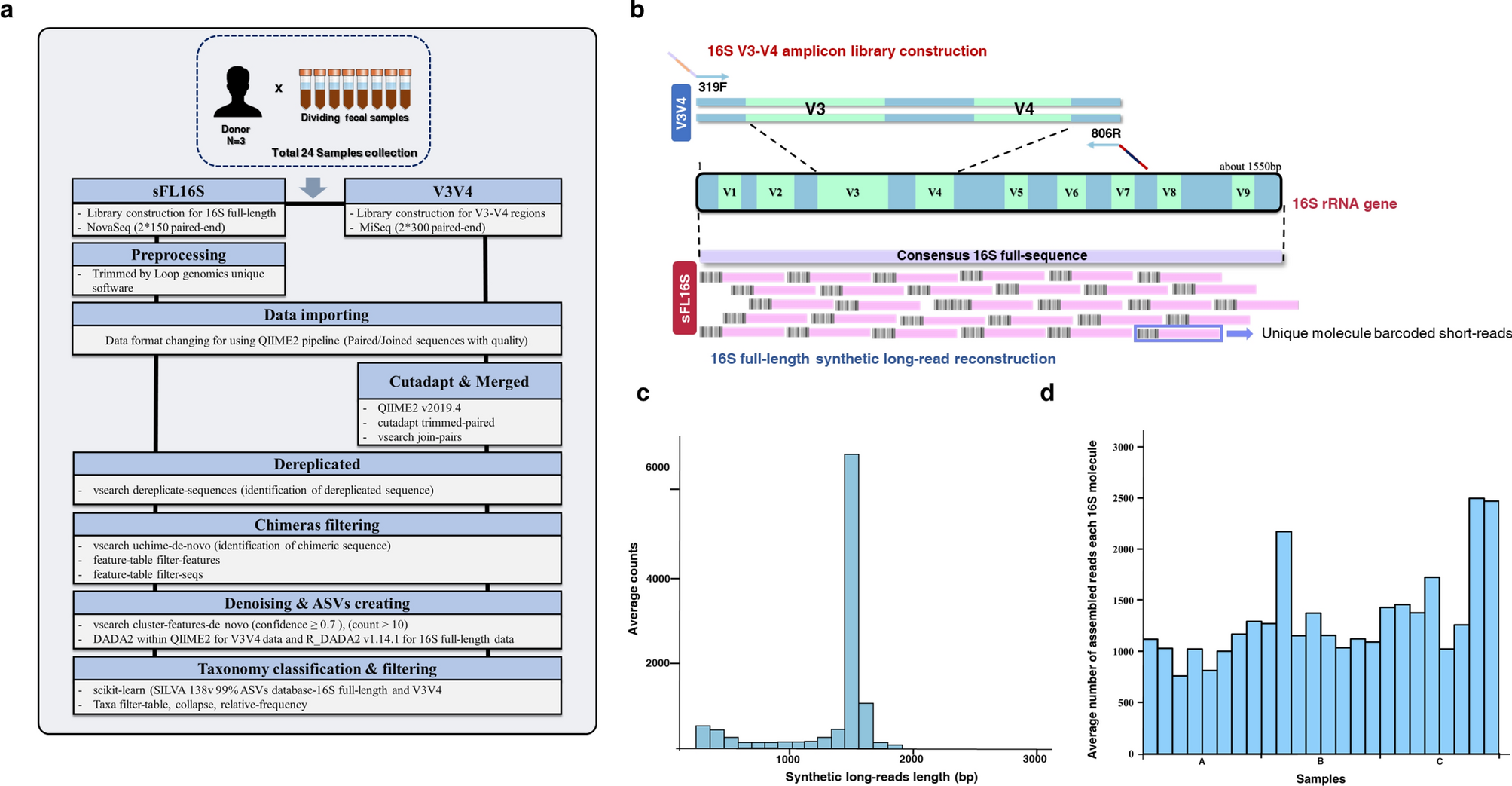
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports

16S rDNA sequence analysis of the three water samples using the SILVA... | Download Scientific Diagram
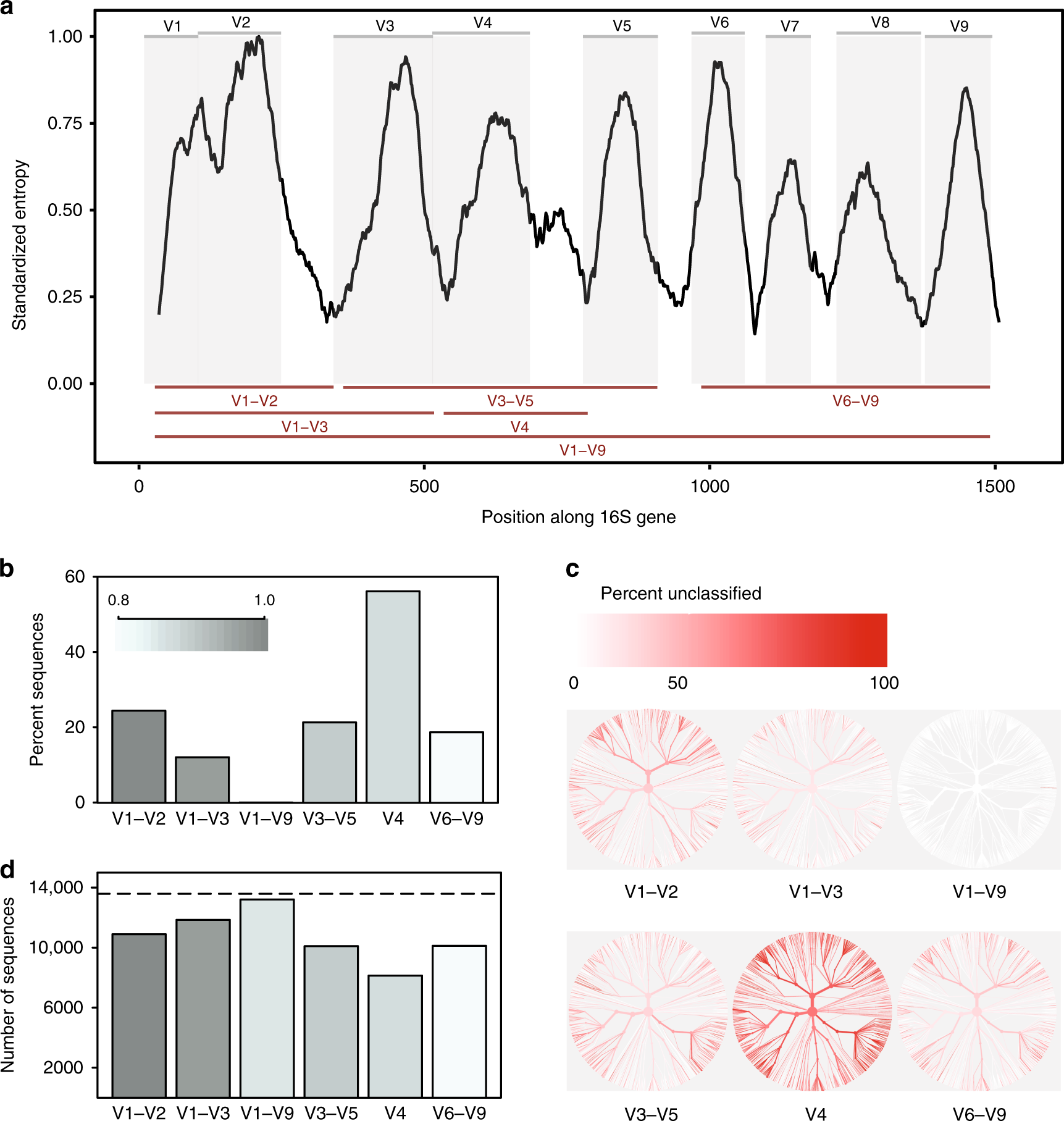
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis | Nature Communications

25 years of serving the community with ribosomal RNA gene reference databases and tools - ScienceDirect
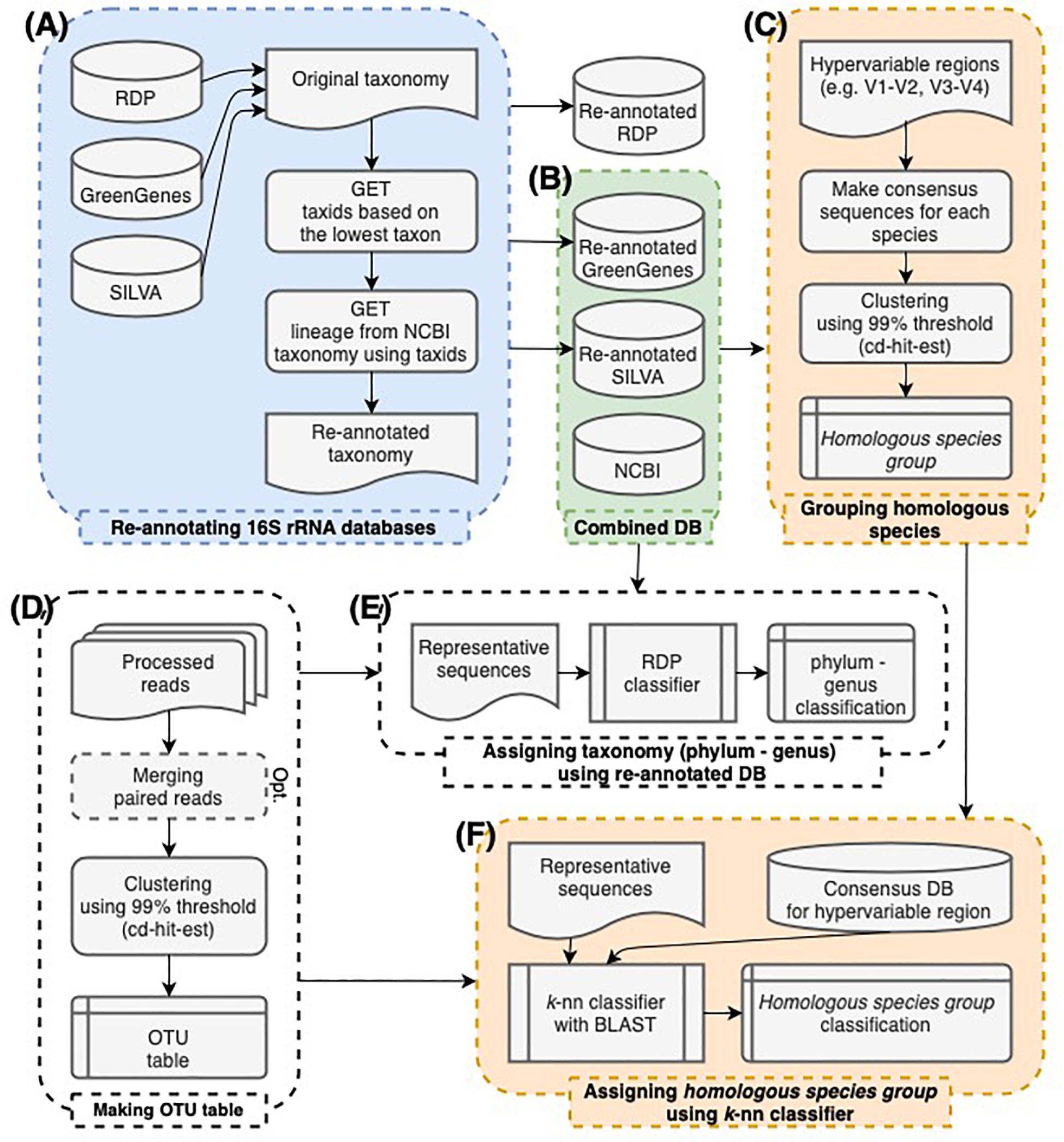
Frontiers | Data-Driven Modeling for Species-Level Taxonomic Assignment From 16S rRNA: Application to Human Microbiomes

Cydrasil 3, a curated 16S rRNA gene reference package and web app for cyanobacterial phylogenetic placement | Scientific Data

The SILVA Database Project: An ELIXIR core data resource for high-quality ribosomal RNA sequences | Semantic Scholar
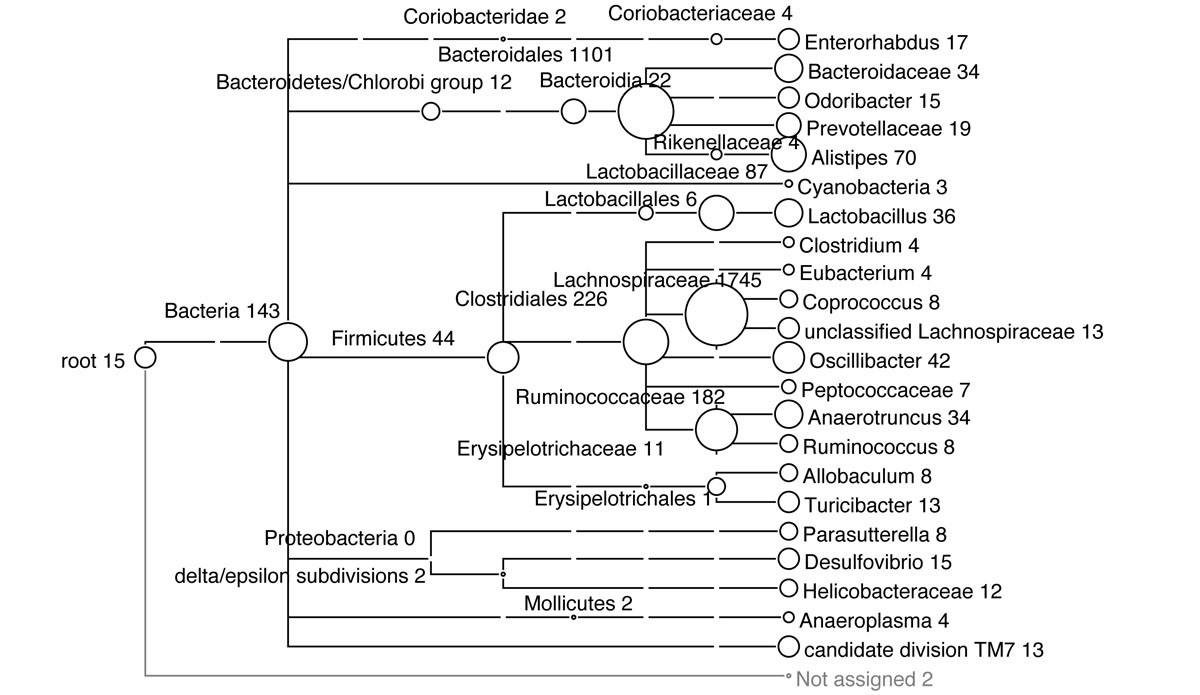
![Improved taxonomic assignment of rumen bacterial 16S rRNA sequences using a revised SILVA taxonomic framework [PeerJ] Improved taxonomic assignment of rumen bacterial 16S rRNA sequences using a revised SILVA taxonomic framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/6496/1/fig-2-full.png)
Improved taxonomic assignment of rumen bacterial 16S rRNA sequences using a revised SILVA taxonomic framework [PeerJ]
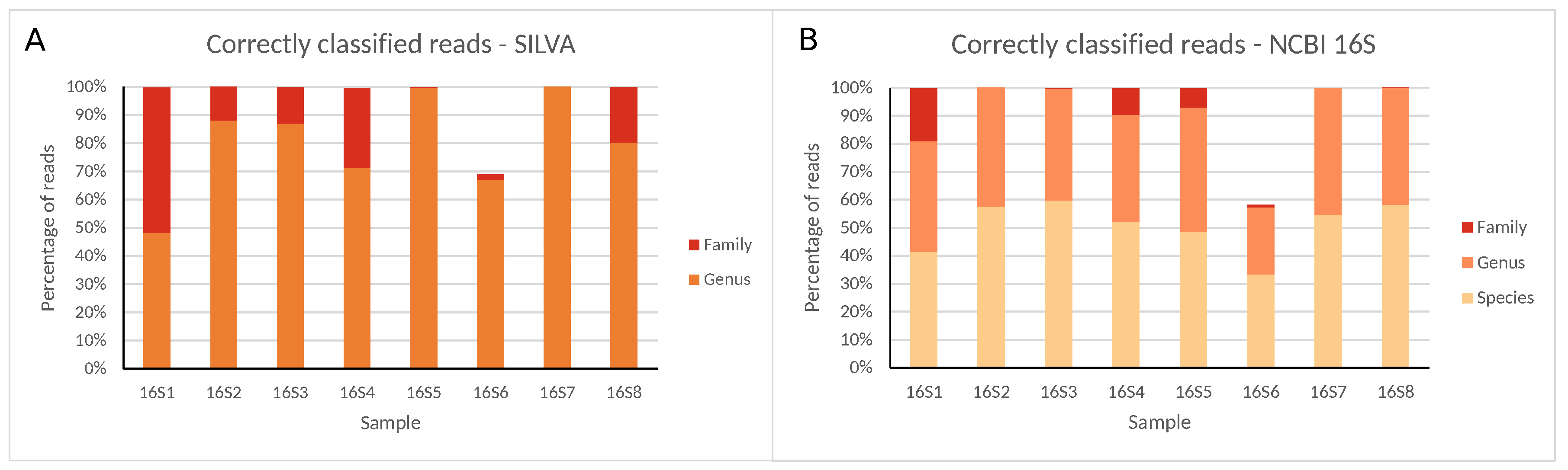
IJMS | Free Full-Text | Targeting the 16S rRNA Gene for Bacterial Identification in Complex Mixed Samples: Comparative Evaluation of Second (Illumina) and Third (Oxford Nanopore Technologies) Generation Sequencing Technologies
GitHub - anweshmaile/silva-138_classifiers: Classifiers trained on different variable regions of Prokaryotic 16S rRNA genes